| country | year | cases | population |
|---|---|---|---|
| Afghanistan | 1999 | 745 | 19987071 |
| Afghanistan | 2000 | 2666 | 20595360 |
| Brazil | 1999 | 37737 | 172006362 |
| Brazil | 2000 | 80488 | 174504898 |
| China | 1999 | 212258 | 1272915272 |
| China | 2000 | 213766 | 1280428583 |
Data Joins and Transformations Review
This module consists of readings reviewing material typically taught in STAT 331. It is possible you can skip over portions of this reading. It is your responsibility to decide which areas you need to review before diving into Stat 541.
Answer the following questions to see if you can safely skip this section.
Each question focuses on the following data tables:
| country | year | type | count |
|---|---|---|---|
| Afghanistan | 1999 | cases | 745 |
| Afghanistan | 1999 | population | 19987071 |
| Afghanistan | 2000 | cases | 2666 |
| Afghanistan | 2000 | population | 20595360 |
| Brazil | 1999 | cases | 37737 |
| Brazil | 1999 | population | 172006362 |
| Brazil | 2000 | cases | 80488 |
| Brazil | 2000 | population | 174504898 |
| China | 1999 | cases | 212258 |
| China | 1999 | population | 1272915272 |
| China | 2000 | cases | 213766 |
| China | 2000 | population | 1280428583 |
| country | year | rate |
|---|---|---|
| Afghanistan | 1999 | 745/19987071 |
| Afghanistan | 2000 | 2666/20595360 |
| Brazil | 1999 | 37737/172006362 |
| Brazil | 2000 | 80488/174504898 |
| China | 1999 | 212258/1272915272 |
| China | 2000 | 213766/1280428583 |
| country | 1999 | 2000 |
|---|---|---|
| Afghanistan | 745 | 2666 |
| Brazil | 37737 | 80488 |
| China | 212258 | 213766 |
| country | 1999 | 2000 |
|---|---|---|
| Afghanistan | 19987071 | 20595360 |
| Brazil | 172006362 | 174504898 |
| China | 1272915272 | 1280428583 |
| country | century | year | rate |
|---|---|---|---|
| Afghanistan | 19 | 99 | 745/19987071 |
| Afghanistan | 20 | 00 | 2666/20595360 |
| Brazil | 19 | 99 | 37737/172006362 |
| Brazil | 20 | 00 | 80488/174504898 |
| China | 19 | 99 | 212258/1272915272 |
| China | 20 | 00 | 213766/1280428583 |
- For each of the Tables (datasets) above, answer the following question: Which of the principles of tidy data does the table violate?
- Variables are spread across multiple columns
- The same observation occupies multiple rows
- There are multiple values in each cell
If you had a hard time answering these questions, I would recommend reviewing Section 1.1.
- Pivot Table 2 so it instead looks like this:
| country | year | cases | population |
|---|---|---|---|
| Afghanistan | 1999 | 745 | 19987071 |
| Afghanistan | 2000 | 2666 | 20595360 |
| Brazil | 1999 | 37737 | 172006362 |
| Brazil | 2000 | 80488 | 174504898 |
| China | 1999 | 212258 | 1272915272 |
| China | 2000 | 213766 | 1280428583 |
If you had a hard time answering this question, I would recommend reviewing Section 1.2.2.
- Use
separate_wider_delim()to separate theratecolumn of Table 3 so it instead looks like this:
| country | year | cases | population |
|---|---|---|---|
| Afghanistan | 1999 | 745 | 19987071 |
| Afghanistan | 2000 | 2666 | 20595360 |
| Brazil | 1999 | 37737 | 172006362 |
| Brazil | 2000 | 80488 | 174504898 |
| China | 1999 | 212258 | 1272915272 |
| China | 2000 | 213766 | 1280428583 |
If you had a hard time answering this question, I would recommend reviewing Section 1.3.1.
- Use
unite()to join thecenturyandyearcolumns of Table 5 so it instead looks like this:
| country | year | rate |
|---|---|---|
| Afghanistan | 1999 | 745/19987071 |
| Afghanistan | 2000 | 2666/20595360 |
| Brazil | 1999 | 37737/172006362 |
| Brazil | 2000 | 80488/174504898 |
| China | 1999 | 212258/1272915272 |
| China | 2000 | 213766/1280428583 |
If you had a hard time answering this question, I would recommend reviewing Section 1.3.2.
- Pivot Tables 4a and 4b to long format and then join them to obtain this dataframe:
| country | year | cases | population |
|---|---|---|---|
| Afghanistan | 1999 | 745 | 19987071 |
| Afghanistan | 2000 | 2666 | 20595360 |
| Brazil | 1999 | 37737 | 172006362 |
| Brazil | 2000 | 80488 | 174504898 |
| China | 1999 | 212258 | 1272915272 |
| China | 2000 | 213766 | 1280428583 |
If you had a hard time answering this question, I would recommend reviewing Section 1.2.1 and Section 2.1.
Tidying Data with tidyr
You should feel comfortable:
- Understand the differences between “wide” and “long” format data and how to transition between the two structures.
- Determining what data format is necessary to generate a desired plot or statistical model.
Tidy Data
The concept of tidy data is useful for mapping variables from the data set to elements in a graph, specifications of a model, or aggregating to create summaries. However, what is considered to be “tidy data” format for one task, might not be in the correct “tidy data” format for a different task. It is important for you to consider the end goal when restructuring your data.
Reshaping Data
It’s fairly common for data to come in forms which are convenient for either human viewing or data entry. Unfortunately, these forms aren’t necessarily the most friendly for analysis. There are two main “layouts” a dataset can have:
- “wide” format
- “long” format
The image below shows the same dataset presented in each layout.

Sometimes we will be given data that is in one layout and we will need to transform it into a different layout. This animation illustrates the process of going from a wide data layout to a long data layout, and how to go from a long data layout back to a wide data layout. Here, we are using the pivot_wider() and pivot_longer() functions from the tidyr package in R.
Let’s explore each of these formats in more depth
Longer
Table 4a and 4b are currently stored in a wide format, where the years are spread across multiple columns. We can use pivot_longer() to pivot these data to long format (with one column for all years).
pivot_longer(table4a,
cols = `1999`:`2000`,
names_to = "year",
values_to = "cases") # A tibble: 6 × 3
country year cases
<chr> <chr> <dbl>
1 Afghanistan 1999 745
2 Afghanistan 2000 2666
3 Brazil 1999 37737
4 Brazil 2000 80488
5 China 1999 212258
6 China 2000 213766Notice that I’m surrounding 1999 and 2000 in back ticks (` `). I need to do this because a number is not a syntactically correct variable name, so using cols = 1999:2000 would not work!
Also note that the year column is currently considered to be a character data type, so we would likely need to mutate() it before we do our analysis!
Wider
Table 2 is currently stored in a long format, where the case and population information is stored in a single column named type. We can use pivot_wider() to pivot these data to wide format (with one column for each statistic).
pivot_wider(table2,
names_from = type,
values_from = count)# A tibble: 6 × 4
country year cases population
<chr> <dbl> <dbl> <dbl>
1 Afghanistan 1999 745 19987071
2 Afghanistan 2000 2666 20595360
3 Brazil 1999 37737 172006362
4 Brazil 2000 80488 174504898
5 China 1999 212258 1272915272
6 China 2000 213766 1280428583Notice that I’m not using a cols argument for pivot_wider()! This is because pivot_wider() only modifies the columns specified in the names_from and values_from arguments.
Separating and Uniting
In addition to pivoting, tidyr provides tools to accomplish two additional tasks: separating variables into two different columns using separate(), and uniting (combining) two variables into one column using unite().
separate_wider_delim()
Table 3 (and Table 5) has a rate column that includes information about both the cases and the population, separated with a / symbol. We can use separate_wider_delim() to create two separate columns for these variables.
separate_wider_delim(table3,
cols = rate,
delim = "/",
names = c("cases", "population"),
cols_remove = TRUE
) # A tibble: 6 × 4
country year cases population
<chr> <dbl> <chr> <chr>
1 Afghanistan 1999 745 19987071
2 Afghanistan 2000 2666 20595360
3 Brazil 1999 37737 172006362
4 Brazil 2000 80488 174504898
5 China 1999 212258 1272915272
6 China 2000 213766 1280428583Note that we could have also used separate_wider_regex() here to split on the / symbol. The patterns argument of separate_wider_regex() is a bit funky:
separate_wider_regex(table3,
rate,
patterns = c(cases = "\\d+", "/", population = "\\d+")
) patterns needs to be a named character vector where the names become column names and the values are regular expressions that match the contents of the vector. cases comes from the first one or more (+) digits \\d, then the split is made on the \, and then population comes from the following one or more digits.
Personally, we think the delim option is a bit easier, but this is how it would work with regular expressions.
unite()
In addition to having variables that need separating, Table 5 also has two variables (century and year) that need to be united (i.e., joined together). We can perform this action using the unite() function.
# A tibble: 6 × 4
country year cases population
<chr> <chr> <chr> <chr>
1 Afghanistan 1999 745 19987071
2 Afghanistan 2000 2666 20595360
3 Brazil 1999 37737 172006362
4 Brazil 2000 80488 174504898
5 China 1999 212258 1272915272
6 China 2000 213766 1280428583Notice that I’m using the pipe operator (|>) to chain these operations together. First, the century and year variables are united and then the resulting dataframe is fed into the separate_wider_delim() function to separate the rate column.
The pipe operator saves us from having to save intermediate objects and makes the process easier to read!
Joining data with dplyr
You should feel comfortable:
- Finding the keys that match between two relational datasets
- Identifying which join(s) will result in the dataset you want
- Using filtering joins in place of the
filter()function
There are two main types of table joins:
Mutating joins, which add columns from one table to matching rows in another table
Filtering joins, which remove rows from a table based on whether or not there is a matching row in another table (but the columns in the original table don’t change)
Mutating Joins
We’re primarily going to focus on mutating joins, as filtering joins can be accomplished by…filtering…rather than by table joins.
An inner_join() only keeps any rows that are included in both tables, but keeps every column.
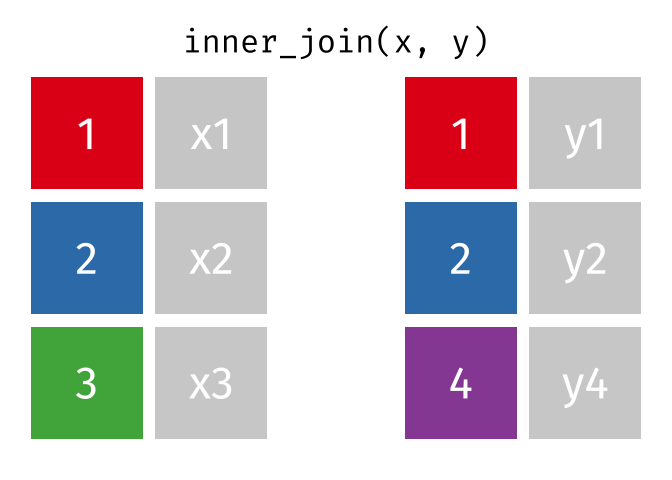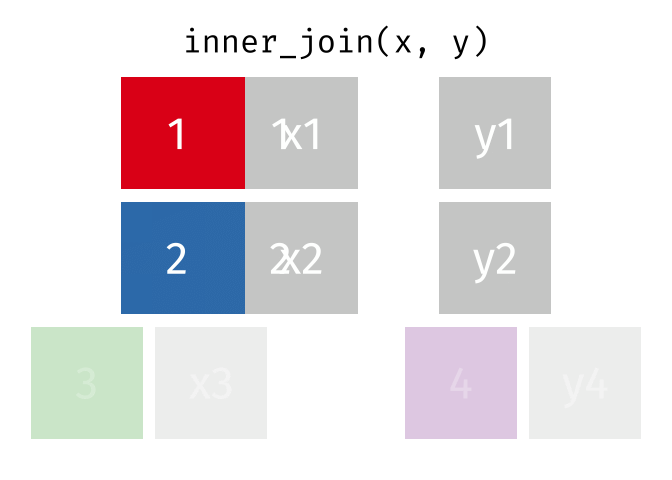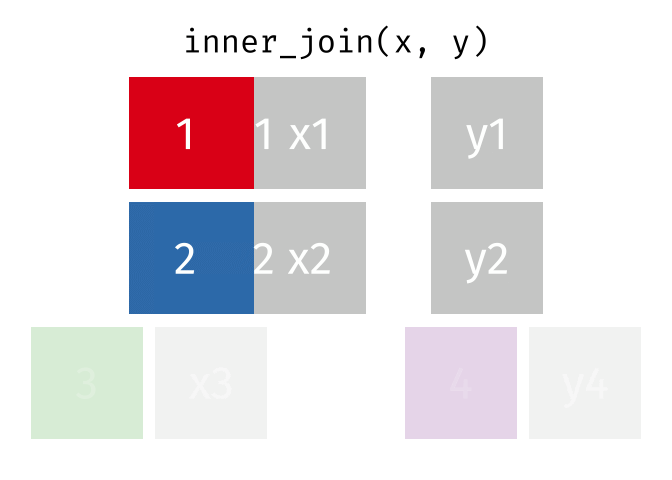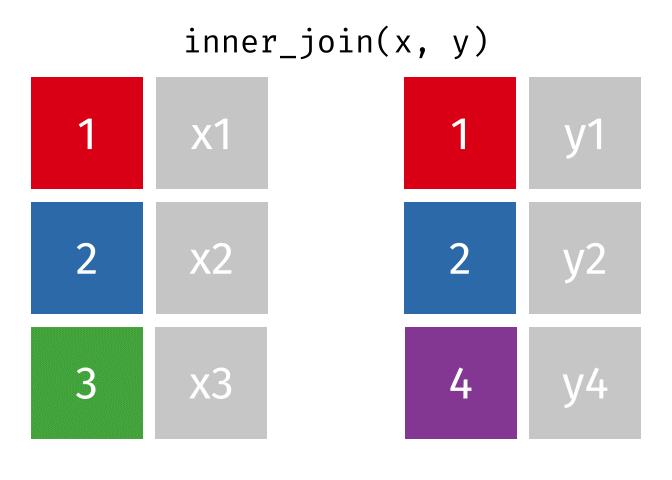
But what if we want to keep all of the rows in x? We would do a left_join() (since x is on the left).
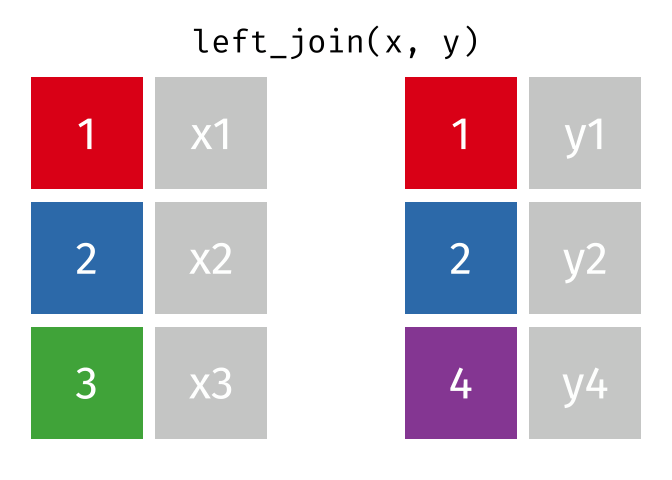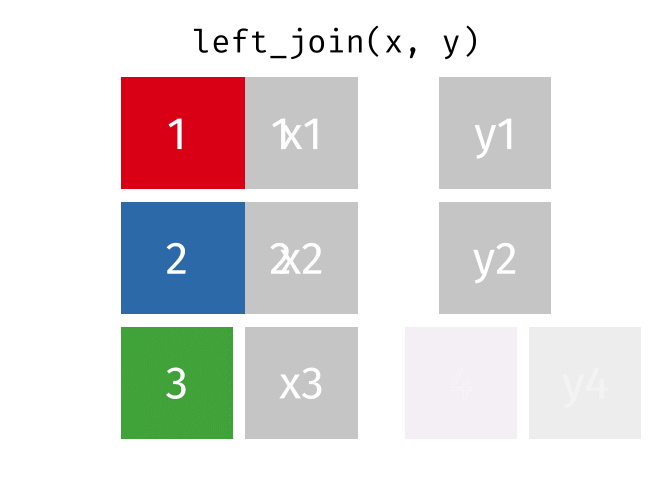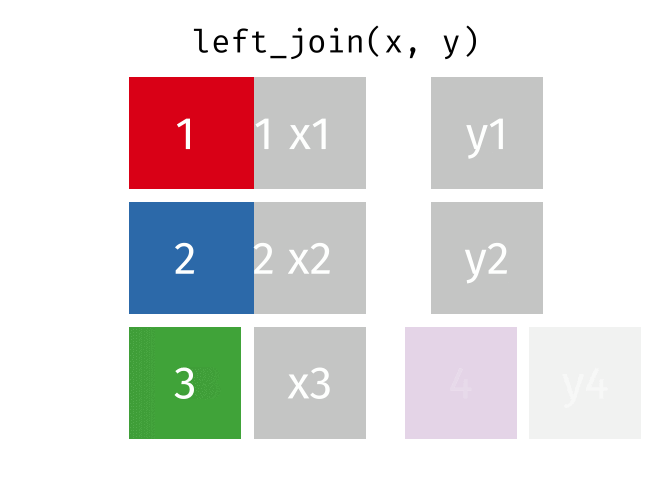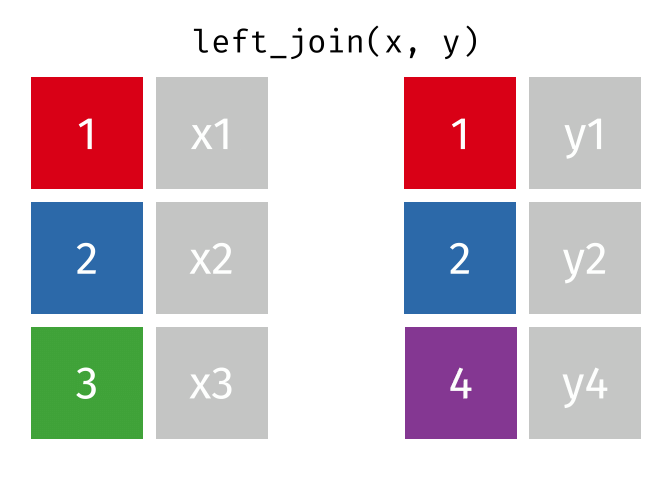
If there are multiple matches in the y table, though, we might have to duplicate rows in x. This is still a left join, just a more complicated one.

If we wanted to keep all of the rows in y, we would do a right_join() (since y is on the right):

We can always change a left join to a right join or visa versa. People seem to have a preference for left joins, but if you like right joins more go for it!
And finally, if we want to keep all of the rows of both x and y, we’d do a full_join():

Every join has a “left side” and a “right side” - so in XXXX_join(A, B), A is the left side, and B is the right side.
Joins are differentiated based on how they treat the rows and columns of each side. In mutating joins, the columns from both sides are always kept. The table below summarizes which rows are kept from each side of the join, for different mutating joins.
| Join Type | Left Side | Right Side |
|---|---|---|
| inner | matching | matching |
| left | all | matching |
| right | matching | all |
| outer | all | all |
Filtering Joins
These joins do not merge two datasets together! Instead, they filter the values of one dataset based on the values of another dataset.
A semi_join() keeps rows in x that have a match in y. Notice that the resulting dataset is a modification of the original x dataset—nothing from y has been added to x!
You can think of this as a relative of using the inclusion operator (%in%) inside of a filter() statement (e.g., filter(day %in% c("Mon", "Tues", "Wed"))).

Suppose we wanted to only keep countries in Table 2 that were also present in Table 1. We could do this using a semi-join:
semi_join(x = table2,
y = table1,
by = "country")# A tibble: 12 × 4
country year type count
<chr> <dbl> <chr> <dbl>
1 Afghanistan 1999 cases 745
2 Afghanistan 1999 population 19987071
3 Afghanistan 2000 cases 2666
4 Afghanistan 2000 population 20595360
5 Brazil 1999 cases 37737
6 Brazil 1999 population 172006362
7 Brazil 2000 cases 80488
8 Brazil 2000 population 174504898
9 China 1999 cases 212258
10 China 1999 population 1272915272
11 China 2000 cases 213766
12 China 2000 population 1280428583Notice that since Table 2 and Table 1 both contain the same countries the resulting dataset (where the matches were kept) is the same as the original Table 2.
A anti_join() keeps rows in x that do not have a match in y. Notice that the resulting dataset is a modification of the original x dataset—nothing from y has been added to x!
You can think of this as a relative of the filter_out() function, which is a better tool for negating a filter statement (e.g., filter(! day %in% c("Mon", "Tues", "Wed"))).

Suppose we wanted to filter out countries in Table 2 that were also present in Table 1. We could do this using an anti-join:
anti_join(x = table2,
y = table1,
by = "country")# A tibble: 0 × 4
# ℹ 4 variables: country <chr>, year <dbl>, type <chr>, count <dbl>Notice that since Table 2 and Table 1 both contain the same countries the resulting dataset (where the matches were filtered out) is empty.
